Coordonnées
 | DR2 - PhD - HDR Email : gwenaelle.andre@inrae.fr Adress : INRAE - Unité MaIAGE, bât 233, Domaine de Vilvert, 78352 Jouy-en-Josas Cedex Phone : +33 1 34 65 22 64 |
I work as a structural computational biologist at Inra, on the Campus of Jouy-en-Josas, in MaIAGE: Applied Mathematics and Computer Science, from Genome to Environment. This lab is headed by Dr. S. Schbath, and has a co-sharing divisions of MIA (Applied Mathematics and Informatics) and MICA (Microbiology and Food Chain). I belong to the StatInfoMics team headed by Dr. P. Nicolas. My interests deal with computational screening of whole genomes and metagenomes to infer proteins structures and functional annotations. I have special focus on proteins of the gut microbiome which are involved in biomedical properties such as host-pathogen or anti-inflammatory signaling.
My primary interest deals with prokaryotic signaling through phosphorylation and dephosphorylation events, mediated by protein Ser/Thr kinases and protein Tyr phosphatases, respectively. I also have fruitful collaborations with biologists from Micalis-INRA, IPBS Toulouse or collaborators from South America and I aim to infer on the combo sequence/structure/function/dynamics for the analysis of their favorite protein(s).
For all my projects, i combine structural and computational bioinformatics, molecular modeling expertise, X-ray crystallography skills, with in vitro/in vivo data from my collaborators, whose methods span biophysics, biochemistry, cellular biology, neurobiology, and immunology.
Follow me on ResearchGate or Linkedin
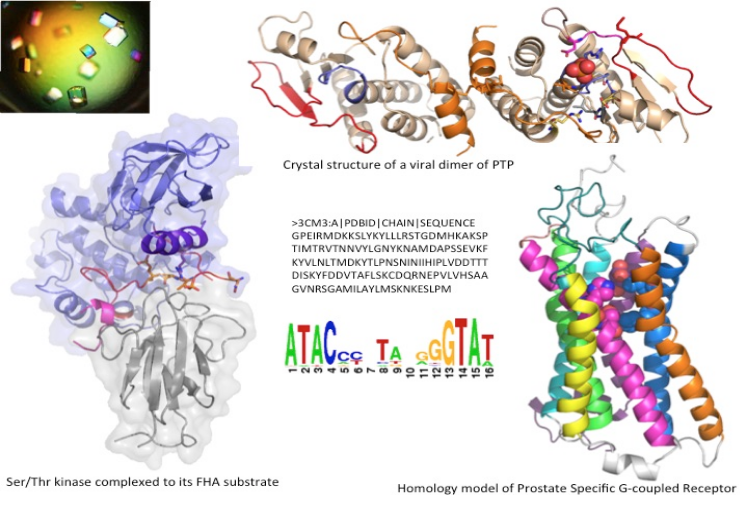
Main collaborations
Faculdad de Ciencias, Montevideo, Uruguay. Coll. A. Villarino & M. Berois. INRAE MaIAGE M. Mariadassou. Projet Ecos-Sud Uruguay 2015: Characterization of the Tyrosine -phosphatase of the intracellular pathogen ORF-virus.
Unidad de Cristalografía de Proteínas. Instituto de Biología Molecular y Celular de Rosario IBR-CONICET-UNR, Argentina. M-N. Lisa. In silico characterization of the conservation of a signaling pathway mediated by PknG in Actinobacteria.
Institut Pasteur, Structural Microbiology Unit, Dir. Prof. P. Alzari. Coll. P. Alzari. Structural and functional characterization of PknB, essential Ser/Thr protein kinase of Mycobacterium tuberculosis.
Institut de Pharmacologie et de Biologie Structurale, Coll. H. Marrakchi M. Daffé. Toulouse. Structural and functional characterization of enzymes responsible for the biogenesis and degradation of mycolic acids in M. tuberculosis.
Institut Pasteur, Human genetics and Cognitive Functions Unit, Dir. Prof. T. Bourgeron. Coll. I. Cloez-Tayarani. Towards structural insights on CNTN6 and Shank3.
INRA Micalis PIMS. Dr. N. Rama Rao, COMBAC Dr. V. Monnet, ProbiHôte Dr. JM. Chatel & P. Langella.
Coordination projects
@2019-2021 : Coordinator of ECOS-Sud project: Structural and functional characterization of Mfd/UvrA interaction.
@2019-2021 : Coordinator in partnership with N. Rama Rao of a TWB project
@2015-2018 : Coordinator of MEM INRA MetaFoldScan
@2015-2017 : Coordinator of ECOS-Sud project: Structural and functional characterization of DUSP Orf virus.
Background
- 2012 - Today : Senior Scientist -structuralist- at INRAE Jouy-en-Josas, MaIAGE SIO.
- 2004-2012 : Senior scientist -structuralist- detached at Institut Pasteur Paris, Structural Microbiology Unit. Skills in X-ray crystallography.
- 2000-2004 : Junior scientist INRA Nantes, BIA. Structural bioinformatics.
- 1999-2000 : Cornell University, Ithaca, USA. Post-doc in Molecular dynamics J.W. Brady's lab at Food Science
- 1994-1998 : INRA, Nantes. PhD in Molecular Mechanics and modeling.
